Leaf of the binary tree. The chemPoint stores the composition 'phi', the mapping of this composition Rphi, the mapping gradient matrix A and the matrix describing the Ellipsoid Of Accuracy (EOA). More...
Public Member Functions | |
| chemPointISAT (chemistryTabulationMethods::ISAT &table, const scalarField &phi, const scalarField &Rphi, const scalarSquareMatrix &A, const scalarField &scaleFactor, const scalar tolerance, const label completeSpaceSize, const label nActive, const dictionary &coeffsDict, binaryNode *node=nullptr) | |
| Construct from components. More... | |
| chemPointISAT (const chemPointISAT &p, binaryNode *node) | |
| Construct from another chemPoint and reference to a binary node. More... | |
| chemPointISAT (chemPointISAT &p) | |
| Construct from another chemPoint. More... | |
| const chemistryTabulationMethods::ISAT & | table () const |
| Access to the ISAT table. More... | |
| label | nGrowth () const |
| label & | completeSpaceSize () |
| const label & | completeSpaceSize () const |
| const scalarField & | phi () const |
| const scalarField & | Rphi () const |
| const scalarField & | scaleFactor () const |
| const scalar & | tolerance () const |
| binaryNode *& | node () |
| const scalarSquareMatrix & | A () const |
| scalarSquareMatrix & | A () |
| const scalarSquareMatrix & | LT () const |
| scalarSquareMatrix & | LT () |
| label | nActive () const |
| const List< label > & | completeToSimplifiedIndex () const |
| const List< label > & | simplifiedToCompleteIndex () const |
| void | increaseNumRetrieve () |
| Increases the number of retrieves the chempoint has generated. More... | |
| void | resetNumRetrieve () |
| Resets the number of retrieves at each time step. More... | |
| void | increaseNLifeTime () |
| Increases the "counter" of the chP life. More... | |
| label | simplifiedToCompleteIndex (const label i) |
| const label & | timeTag () |
| label & | lastTimeUsed () |
| bool & | toRemove () |
| label & | maxNumNewDim () |
| const label & | numRetrieve () |
| const label & | nLifeTime () |
| bool | inEOA (const scalarField &phiq) |
| To RETRIEVE the mapping from the stored chemPoint phi, the query. More... | |
| bool | grow (const scalarField &phiq) |
| More details about the minimum-volume ellipsoid covering an. More... | |
| bool | checkSolution (const scalarField &phiq, const scalarField &Rphiq) |
| If phiq is not in the EOA, then the mapping is computed. More... | |
Static Public Member Functions | |
| static void | changeTolerance (scalar newTol) |
Leaf of the binary tree. The chemPoint stores the composition 'phi', the mapping of this composition Rphi, the mapping gradient matrix A and the matrix describing the Ellipsoid Of Accuracy (EOA).
1) When the chemPoint is created the region of accuracy is approximated by an ellipsoid E centered in 'phi' (obtained with the constant): E = {x| ||L^T.(x-phi)|| <= 1}, with x a point in the composition space and L^T the transpose of an upper triangular matrix describing the EOA (see below: "Computation of L" ).
2) To RETRIEVE the mapping from the chemPoint phi, the query point phiq has to be in the EOA of phi. It follows that, dphi=phiq-phi and to test if phiq is in the ellipsoid there are two methods. First, compare r=||dphi|| with rmin and rmax. If r < rmin, phiq is in the EOA. If r > rmax, phiq is out of the EOA. This operations is O(completeSpaceSize) and is performed first. If rmin < r < rmax, then the second method is used: ||L^T.dphi|| <= 1 to be in the EOA. If phiq is in the EOA, Rphiq is obtained by linear interpolation: Rphiq= Rphi + A.dphi.
3) If phiq is not in the EOA, then the mapping is computed. But as the EOA is a conservative approximation of the region of accuracy surrounding the point phi, we could expand it by comparing the computed results with the one obtained by linear interpolation. The error epsGrow is calculated: epsGrow = ||B.(dR - dRl)||, with dR = Rphiq - Rphi, dRl = A.dphi and B the diagonal scale factor matrix. If epsGrow <= tolerance, the EOA is too conservative and a GROW is performed otherwise, the newly computed mapping is associated to the initial composition and added to the tree.
4) To GROW the EOA, we expand it to include the previous EOA and the query point phiq. The rank-one matrix method is used. The EOA is transformed to a hypersphere centered at the origin. Then it is expanded to include the transformed point phiq' on its boundary. Then the inverse transformation give the modified matrix L' (see below: "Grow the EOA").
Computation of L : In [1], the EOA of the constant approximation is given by E = {x| ||B.A/tolerance.(x-phi)|| <= 1},
with B a scale factor diagonal matrix, A the mapping gradient matrix and tolerance the absolute tolerance. If we take the QR decomposition of (B.A)/tolerance= Q.R, with Q an orthogonal matrix and R an upper triangular matrix such that the EOA is described by (phiq-phi0)^T.R^T.R.(phiq-phi0) <= 1 L^T = R, both Cholesky decomposition of A^T.B^T.B.A/tolerance^2 This representation of the ellipsoid is used in [2] and in order to avoid large value of semi-axe length in certain direction, a Singular Value Decomposition (SVD) is performed on the L matrix: L = UDV^T, with the orthogonal matrix U giving the directions of the principal axes and 1/di the inverse of the element of the diagonal matrix D giving the length of the principal semi-axes. To avoid very large value of those length, di' = max(di, 1/(alphaEOA*sqrt(tolerance))), with alphaEOA = 0.1 (see [2]) di' = max(di, 1/2), see [1]. The latter will be used in this implementation. And L' = UD'V^T, with D' the diagonal matrix with the modified di'.
Grow the EOA : More details about the minimum-volume ellipsoid covering an ellipsoid E and a point p are found in [3]. Here is the main steps to obtain the modified matrix L' describing the new ellipsoid.
1) calculate the point p' in the transformed space : p' = L^T.(p-phi) 2) compute the rank-one decomposition: G = I + gamma.p'.p'^T, with gamma = (1/|p'|-1)*1/|p'|^2 3) compute L': L' = L.G.
References:
[1] Pope, S. B. (1997).
Computationally efficient implementation of combustion chemistry using
in situ adaptive tabulation.
Combustion Theory and Modelling, 1, 41-63.
[2] Lu, L., & Pope, S. B. (2009).
An improved algorithm for in situ adaptive tabulation.
Journal of Computational Physics, 228(2), 361-386.
[3] Pope, S. B. (2008).
Algorithms for ellipsoids.
Cornell University Report No. FDA, 08-01.
Definition at line 147 of file chemPointISAT.H.
| chemPointISAT | ( | chemistryTabulationMethods::ISAT & | table, |
| const scalarField & | phi, | ||
| const scalarField & | Rphi, | ||
| const scalarSquareMatrix & | A, | ||
| const scalarField & | scaleFactor, | ||
| const scalar | tolerance, | ||
| const label | completeSpaceSize, | ||
| const label | nActive, | ||
| const dictionary & | coeffsDict, | ||
| binaryNode * | node = nullptr |
||
| ) |
Construct from components.
Definition at line 199 of file chemPointISAT.C.
References chemPointISAT::A(), ISAT::chemistry(), chemPointISAT::completeSpaceSize(), odeChemistryModel::cTos(), D, forAll, Foam::max(), Foam::multiply(), ISAT::reduction(), SVD::S(), Foam::fvm::S(), chemPointISAT::scaleFactor(), List< T >::setSize(), chemPointISAT::simplifiedToCompleteIndex(), odeChemistryModel::sToc(), Matrix< Form, Type >::T(), chemPointISAT::tolerance(), SVD::U(), and SVD::V().
| chemPointISAT | ( | const chemPointISAT & | p, |
| binaryNode * | node | ||
| ) |
Construct from another chemPoint and reference to a binary node.
| chemPointISAT | ( | Foam::chemPointISAT & | p | ) |
Construct from another chemPoint.
Definition at line 303 of file chemPointISAT.C.
References chemPointISAT::completeSpaceSize(), and p.

|
inline |
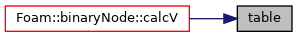
Access to the ISAT table.
Definition at line 27 of file chemPointISATI.H.
Referenced by binaryNode::calcV().

|
inline |
Definition at line 33 of file chemPointISATI.H.
|
inline |
Definition at line 39 of file chemPointISATI.H.
Referenced by binaryNode::calcV(), and chemPointISAT::chemPointISAT().
|
inline |
Definition at line 45 of file chemPointISATI.H.
|
inline |
Definition at line 51 of file chemPointISATI.H.
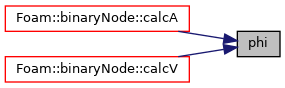
Referenced by binaryNode::calcA(), and binaryNode::calcV().

|
inline |
Definition at line 57 of file chemPointISATI.H.
|
inline |
Definition at line 63 of file chemPointISATI.H.
Referenced by binaryNode::calcV(), and chemPointISAT::chemPointISAT().
|
inline |
Definition at line 69 of file chemPointISATI.H.
Referenced by binaryNode::calcV(), and chemPointISAT::chemPointISAT().

|
inlinestatic |
Definition at line 75 of file chemPointISATI.H.
|
inline |
Definition at line 81 of file chemPointISATI.H.
Referenced by binaryTree::deleteLeaf(), and binaryTree::insertNewLeaf().

|
inline |
Definition at line 87 of file chemPointISATI.H.
Referenced by chemPointISAT::chemPointISAT().

|
inline |
Definition at line 93 of file chemPointISATI.H.
|
inline |
Definition at line 99 of file chemPointISATI.H.
Referenced by binaryNode::calcV().

|
inline |
Definition at line 105 of file chemPointISATI.H.
|
inline |
Definition at line 111 of file chemPointISATI.H.
Referenced by binaryNode::calcV().

|
inline |
Definition at line 118 of file chemPointISATI.H.
Referenced by binaryNode::calcV().

|
inline |
Definition at line 125 of file chemPointISATI.H.
Referenced by chemPointISAT::chemPointISAT().
|
inline |
Increases the number of retrieves the chempoint has generated.
Definition at line 131 of file chemPointISATI.H.
|
inline |
Resets the number of retrieves at each time step.
Definition at line 137 of file chemPointISATI.H.
Referenced by binaryTree::resetNumRetrieve().

|
inline |
Increases the "counter" of the chP life.
Definition at line 143 of file chemPointISATI.H.
|
inline |
Definition at line 149 of file chemPointISATI.H.
|
inline |
Definition at line 177 of file chemPointISATI.H.
|
inline |
Definition at line 183 of file chemPointISATI.H.
|
inline |
Definition at line 189 of file chemPointISATI.H.
|
inline |
Definition at line 195 of file chemPointISATI.H.
|
inline |
Definition at line 201 of file chemPointISATI.H.
|
inline |
Definition at line 207 of file chemPointISATI.H.
| bool inEOA | ( | const scalarField & | phiq | ) |
To RETRIEVE the mapping from the stored chemPoint phi, the query.
point phiq has to be in the EOA of phi. To test if phiq is in the ellipsoid: ||L^T.dphi|| <= 1
Definition at line 337 of file chemPointISAT.C.
References Foam::endl(), Foam::Info, Foam::max(), Foam::sqr(), and Foam::sqrt().
Referenced by binaryTree::secondaryBTSearch().

| bool grow | ( | const scalarField & | phiq | ) |
More details about the minimum-volume ellipsoid covering an.
ellipsoid E and a point p are found in [1]. Here is the main steps to obtain the modified matrix L' describing the new ellipsoid. 1) calculate the point p' in the transformed space : p' = L^T.(p-phi) 2) compute the rank-one decomposition: G = I + gamma.p'.p'^T, with gamma = (1/|p'|-1)*1/|p'|^2 3) compute L': L'L'^T = (L.G)(L.G)^T, L'^T is then obtained by QR decomposition of (L.G)^T = G^T.L^T [1] Stephen B. Pope, "Algorithms for ellipsoids", FDA 08-01, Cornell University, 2008
add new column and line for the new active species transfer last two lines of the previous matrix (p and T) to the end
(change the diagonal position) !set all element of the new lines and columns to zero except diagonal /*! (=1/(tolerance*scaleFactor))
Definition at line 541 of file chemPointISAT.C.
References DynamicList< T, SizeInc, SizeMult, SizeDiv >::append(), forAll, List< T >::size(), Foam::sqr(), Foam::sqrt(), and Foam::Zero.
| bool checkSolution | ( | const scalarField & | phiq, |
| const scalarField & | Rphiq | ||
| ) |
If phiq is not in the EOA, then the mapping is computed.
But as the EOA is a conservative approximation of the region of accuracy surrounding the point phi, we could expand it by comparing the computed results with the one obtained by linear interpolation. The error eps is calculated: eps = ||B.(dR - dRl)||, with dR = Rphiq - Rphi, dRl = A.dphi and B the diagonal scale factor matrix. If eps <= tolerance, the EOA is too conservative and a GROW is performed, otherwise, the newly computed mapping is associated to the initial composition and added to the tree.
Definition at line 476 of file chemPointISAT.C.
References A, Foam::sqr(), and Foam::sqrt().